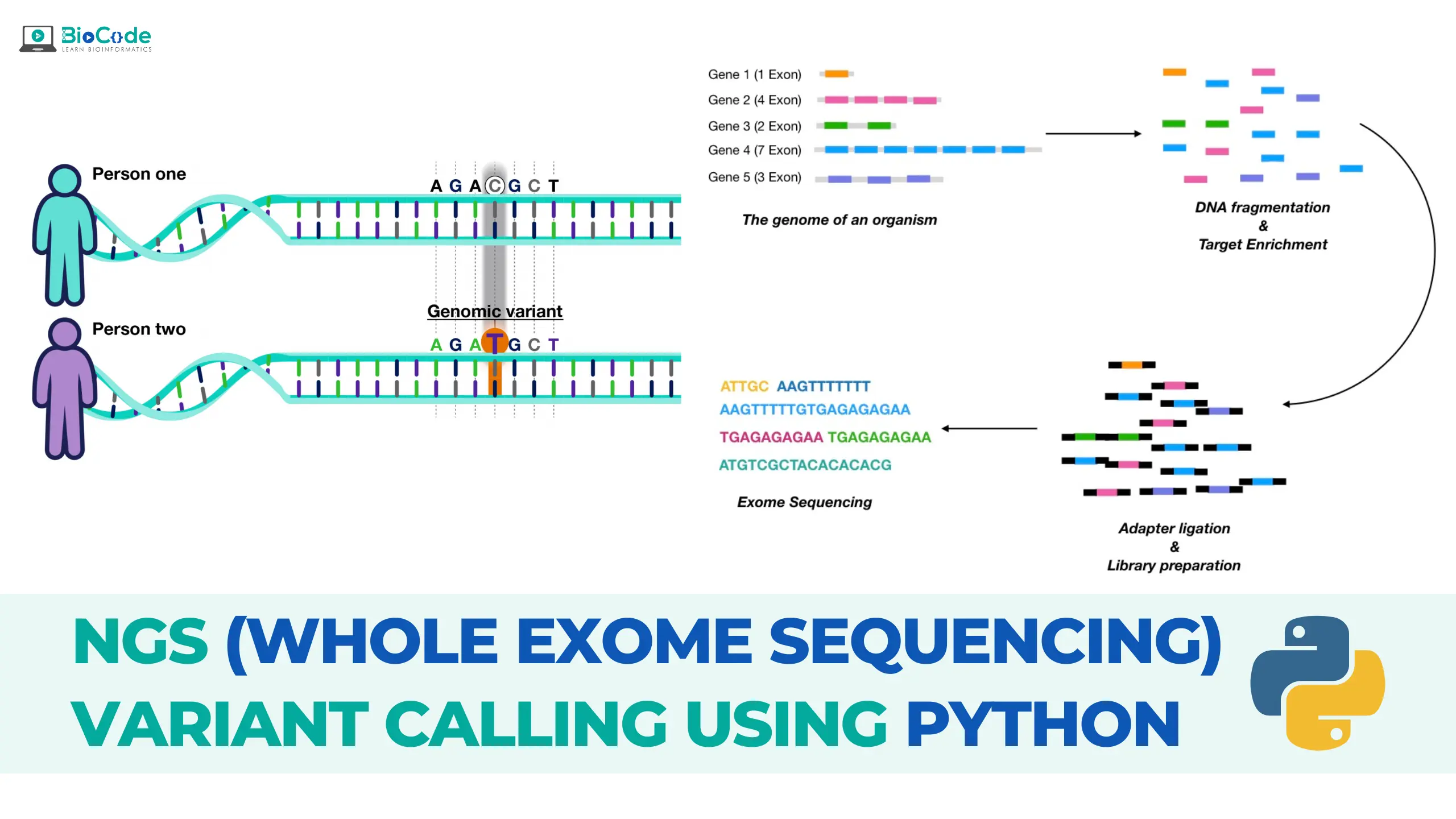
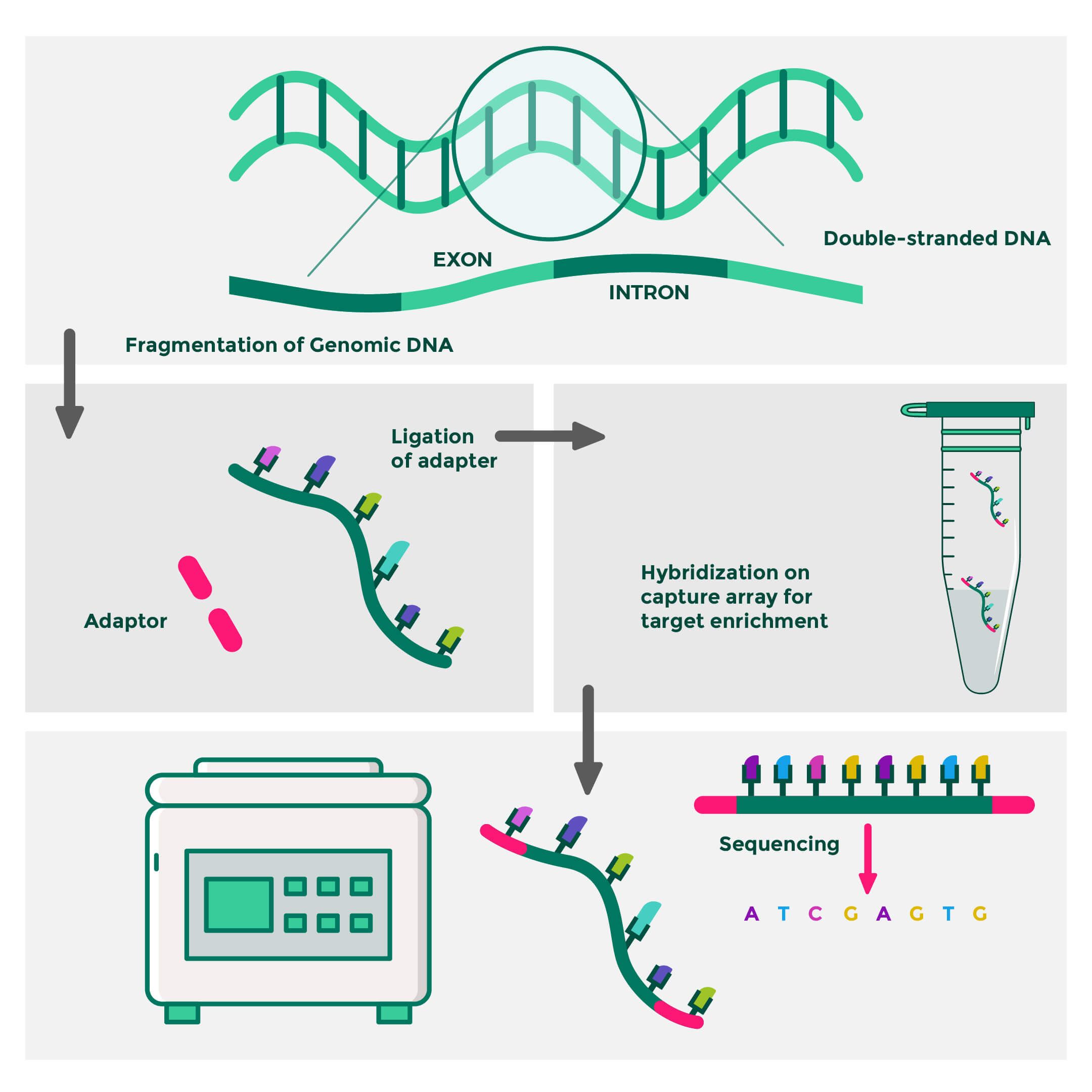
Flowchart of WES (Whole Exome Sequencing) analysis: T1, T2, T3 design... | Download Scientific Diagram
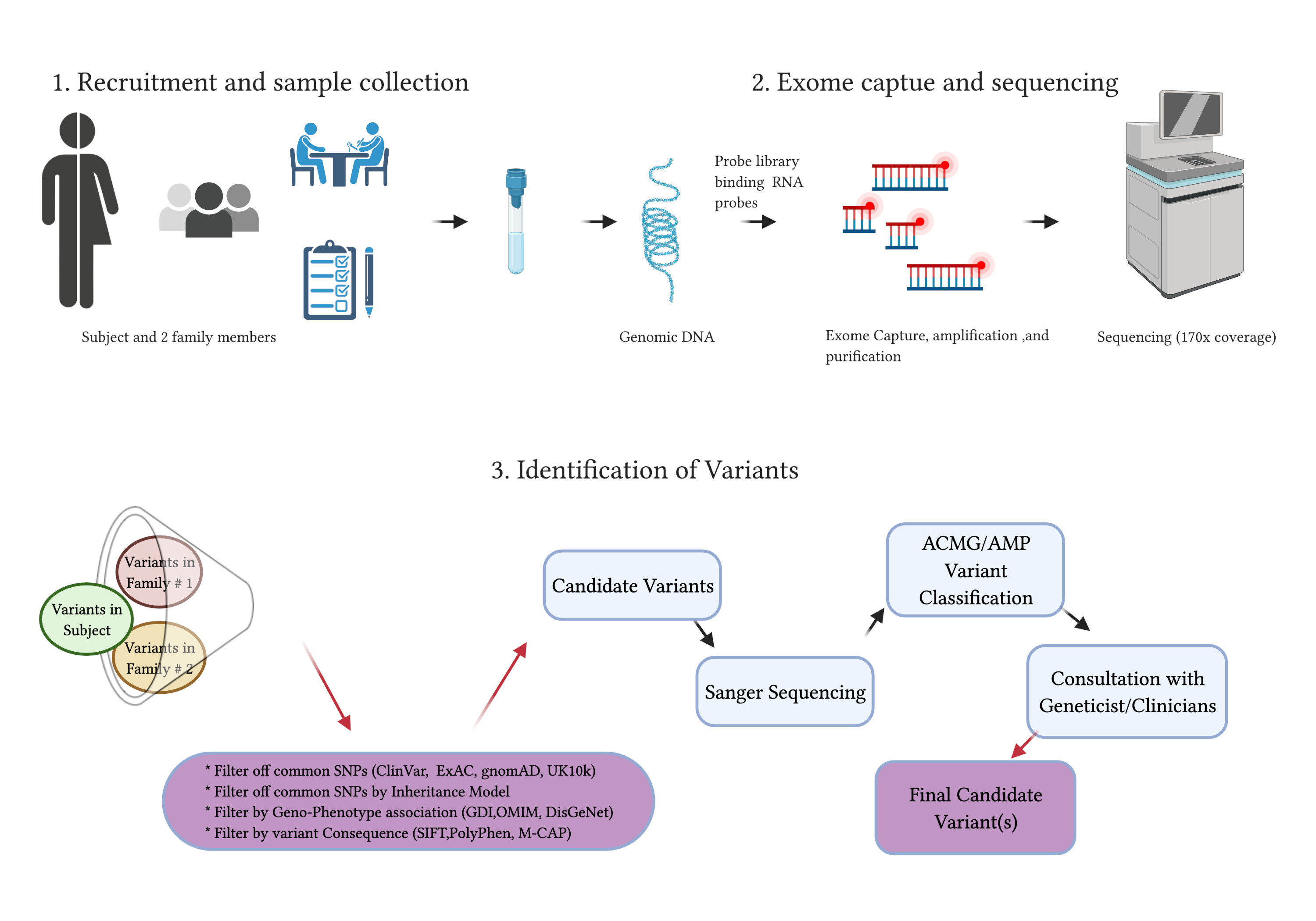
JCM | Free Full-Text | Whole-Exome Sequencing to Identify Potential Genetic Risk in Substance Use Disorders: A Pilot Feasibility Study

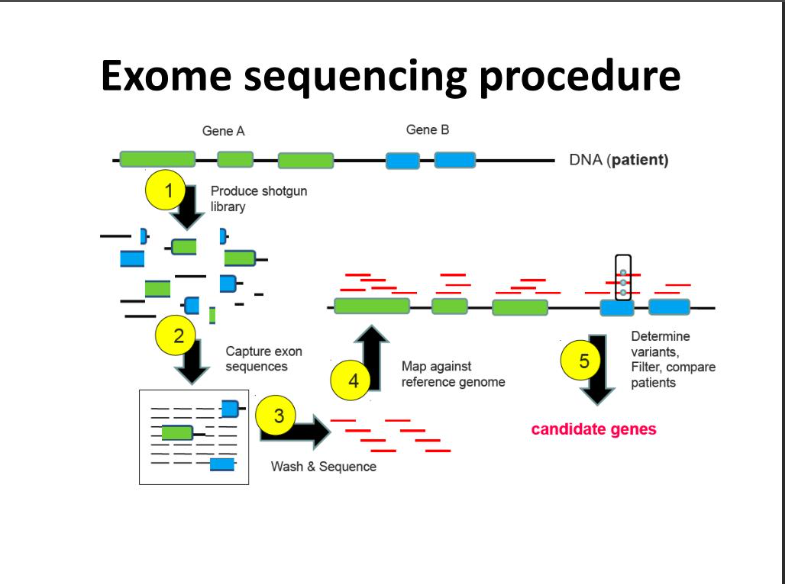
Overview of whole exome sequencing pipeline. SNV, single nucleotide... | Download Scientific Diagram

Accelerated identification of disease-causing variants with ultra-rapid nanopore genome sequencing | Nature Biotechnology
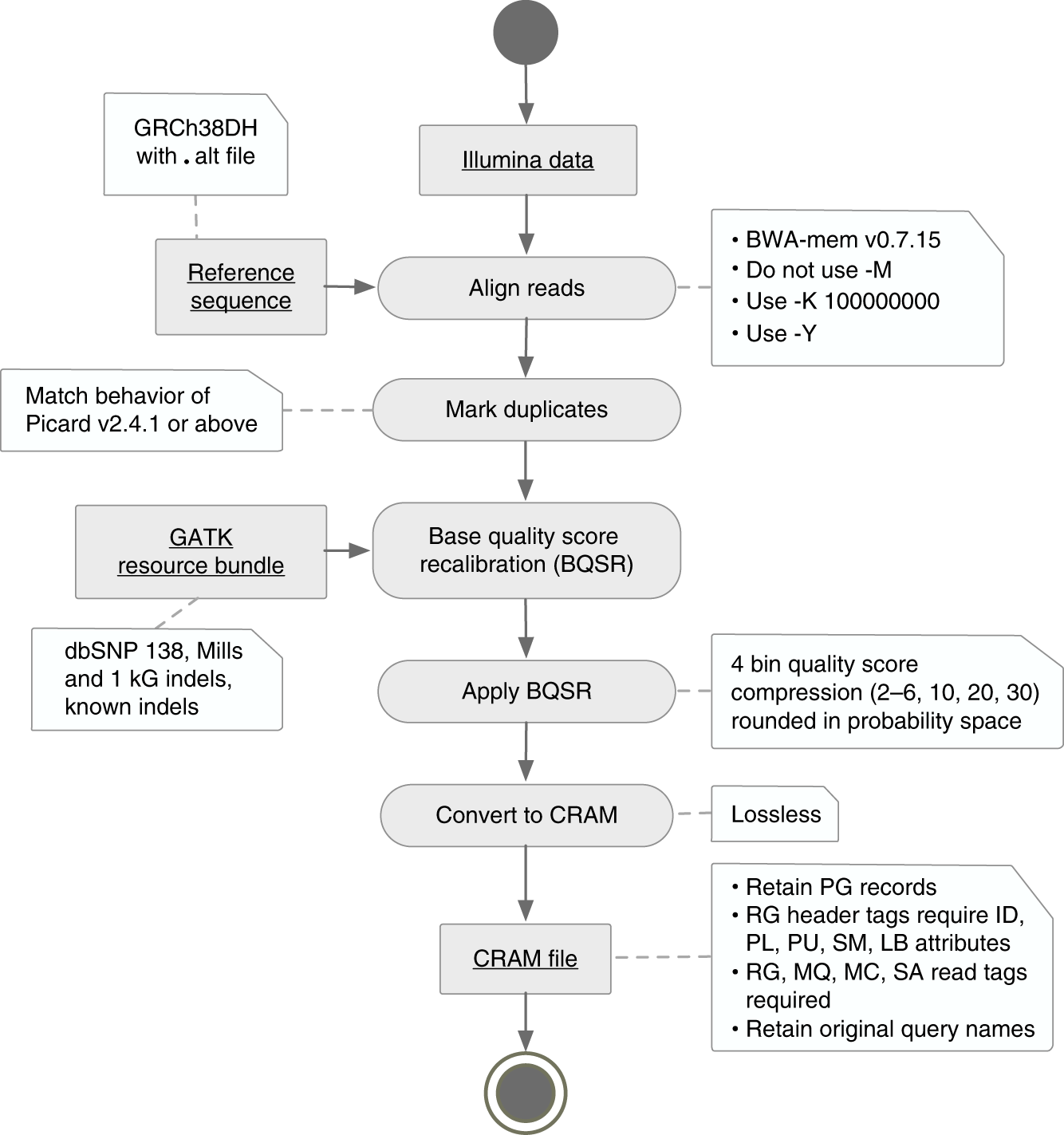
Functional equivalence of genome sequencing analysis pipelines enables harmonized variant calling across human genetics projects | Nature Communications
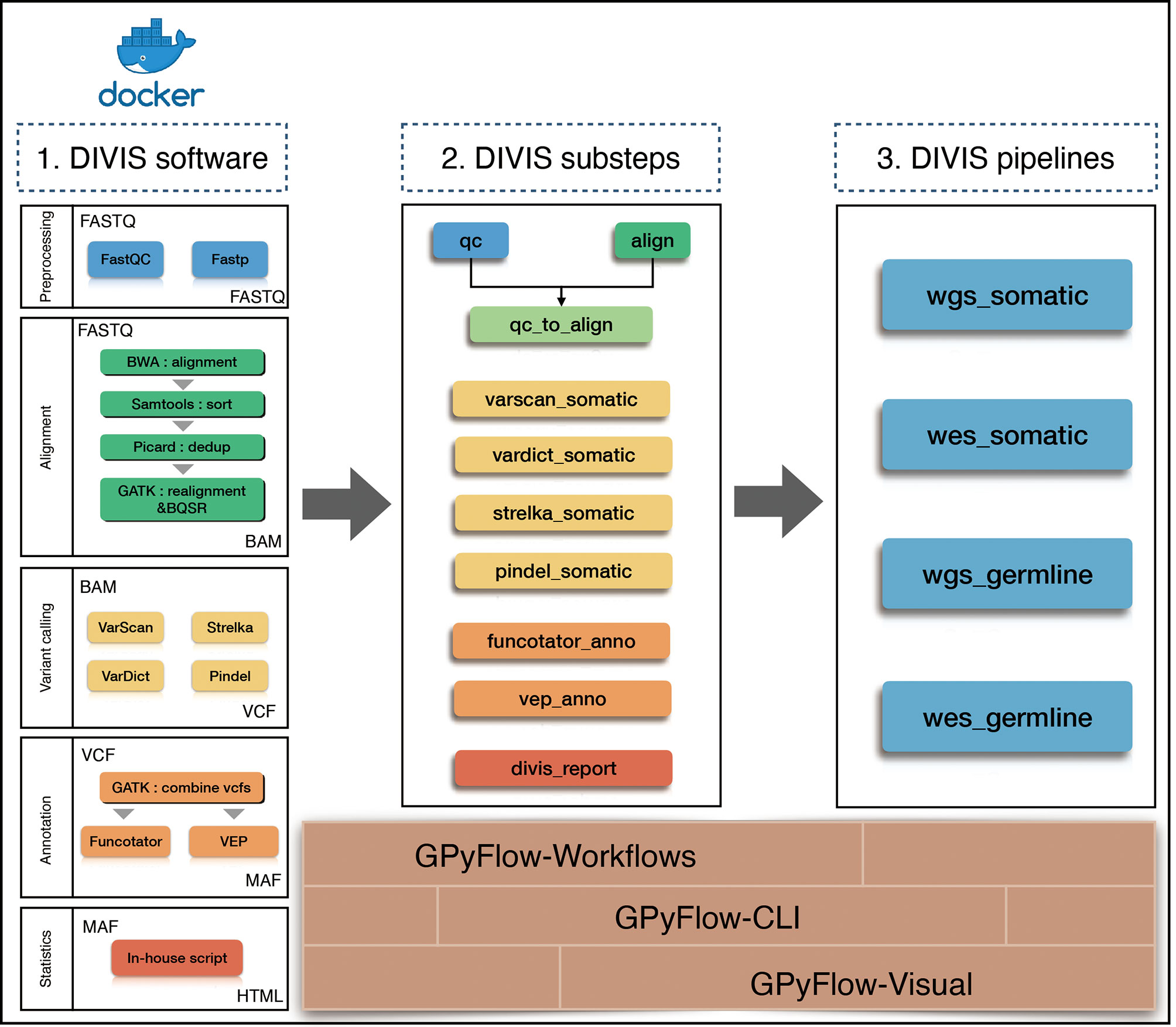
Frontiers | DIVIS: Integrated and Customizable Pipeline for Cancer Genome Sequencing Analysis and Interpretation

Var∣Decrypt: a novel and user-friendly tool to explore and prioritize variants in whole-exome sequencing data | Epigenetics & Chromatin | Full Text